Xun Shi
Dartmouth College
Bottom-up approach to epidemic modeling and access to high-performance computing infrastructure
Epidemic modeling is to build models to represent, characterize, and simulate the spread of an infectious disease in some given population, and ultimately aims to understand and even predict such spread for the purpose of control and intervention of the disease (Andersson and Britton 2000) . Conventional epidemic modeling is a top-down process, i.e., it assumes that the spread of disease can be represented by a generalized mathematical model; specifics of the process can be derived from this model; states of individuals can be inferred based on the state of the population; and situation of local areas is the same as that of the entire area. Conventional epidemic models typically work with aggregate data and focus on the temporal dimension, without considering spatial variation within the study area (Li et al. 2019 ; Li et al. 2020 ).
Individualization and spatialization are considered to be major trends in epidemic modeling, and the two are closely coupled and can naturally converge into one integrated modeling approach (Li et al. 2020). This, on the one hand, is driven by the growing recognition, in the field of epidemiology, of the importance of individual characteristics and spatial variation in epidemic modeling. On the other hand, it is supported and facilitated by the increasing availability of data of patients and the population, the increasing capacity of computing, as well as the rapid development of computational scientific approaches based on that data and capacity (Shi and Wang 2015 ).
Within this context, we have developed a bottom-up epidemic modeling approach, the Epidemic Forest (Li et al. 2019 ). We adapt it to the COVID-19 disease and applied it to real-world combats against the pandemic. The model intends to model the individual-level transmission by taking into account all available information about the patients, the population at risk, and the environment, including but not limited to: epidemiological characteristics, demographics, residence, work, mobility, transportation, social network, vector habitats (in the case of vector-borne diseases), and even genetics. The model represents a disease spread process starting from a primary case using a tree structure. If an epidemic/pandemic has multiple/many primary cases, their epidemic trees form an epidemic forest (herein referred to as epi-forest). The modeling process is to construct this forest based on the modeled individual-level transmission relationship. With the constructed epi-forest, detailed spatiotemporal and epidemiological characteristics of the epidemic/pandemic that may not be available from a top-down model can be quickly and empirically derived, which may provide specific evidence and guidance to planning and decision making of control and intervention, and also inform the prediction and scenario simulation of the epidemic/pandemic.
However, the bottom-up approach to epidemic modeling requires handling large volume of data and performing intensive computation, which may be cumbersome to many researchers and practitioners who do not have easy access to a high-performance computing platform. This difficulty has hindered the adoptability and reproducibility of the approach, which is what we are trying to address with the CyberGIS in this NSF program. With the support of this NSF program, we created a Jupyter Notebook (http://jupyter.org) on the CyberGISX platform (https://cybergisxhub.cigi.illinois.edu/) that provides documents and access to the five modules of the epi-forest modeling process in a sequence, including Disaggregating Trajectory Data, Evaluating Trajectory Overlap, Building Epidemic Forest, Extracting Epidemic Features, and Simulating Scenarios. The current five modules are designed to model a human-human communicable disease like COVID-19. They can also be adapted to suit a vector-borne disease like Dengue Fever or Lyme Disease. The users can choose to run certain modules based on the data they possess and/or the purpose of their project. With the simple register and login process of this platform, a user can easily access this Notebook, try it with the provided test data, and run it with their own data.
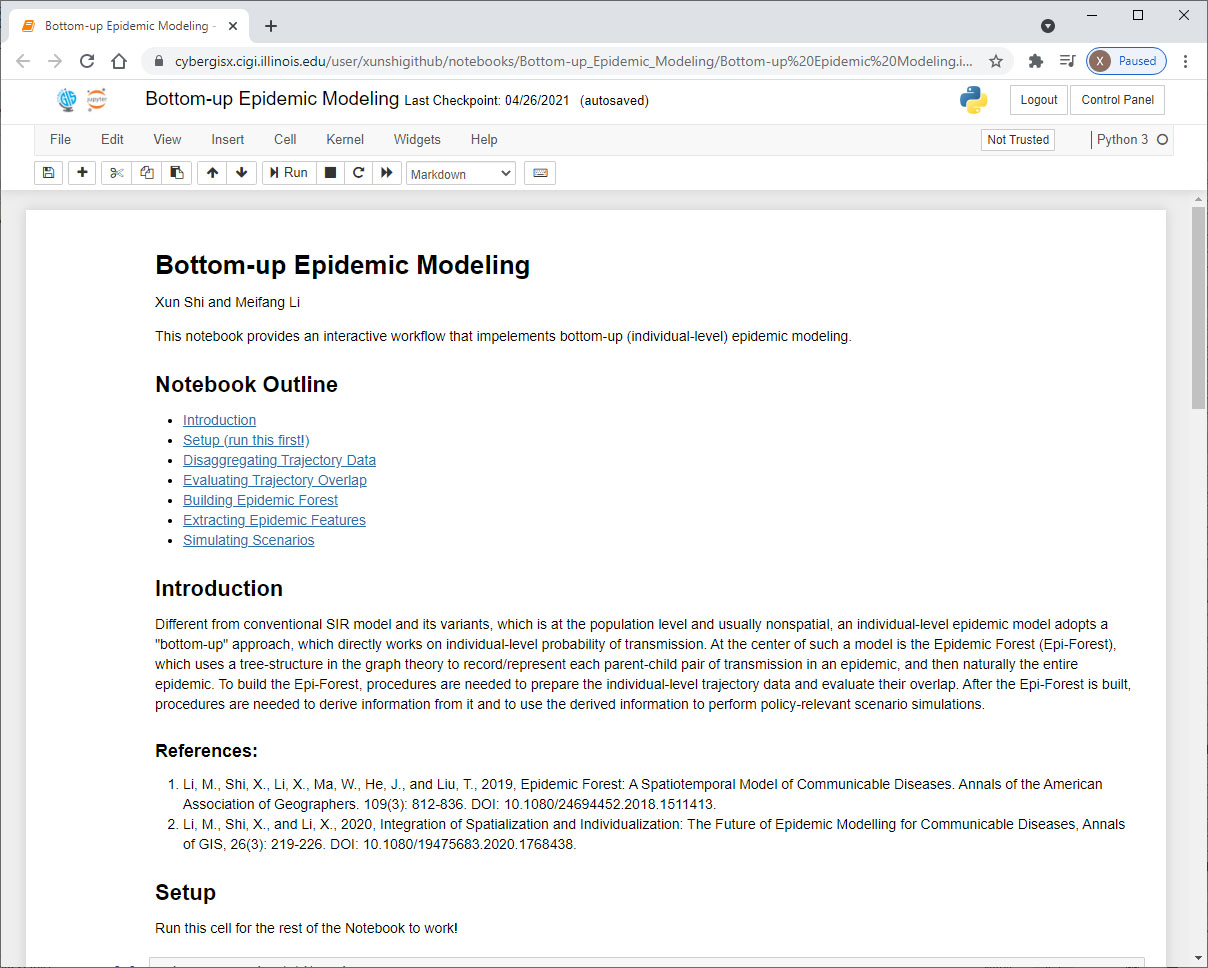
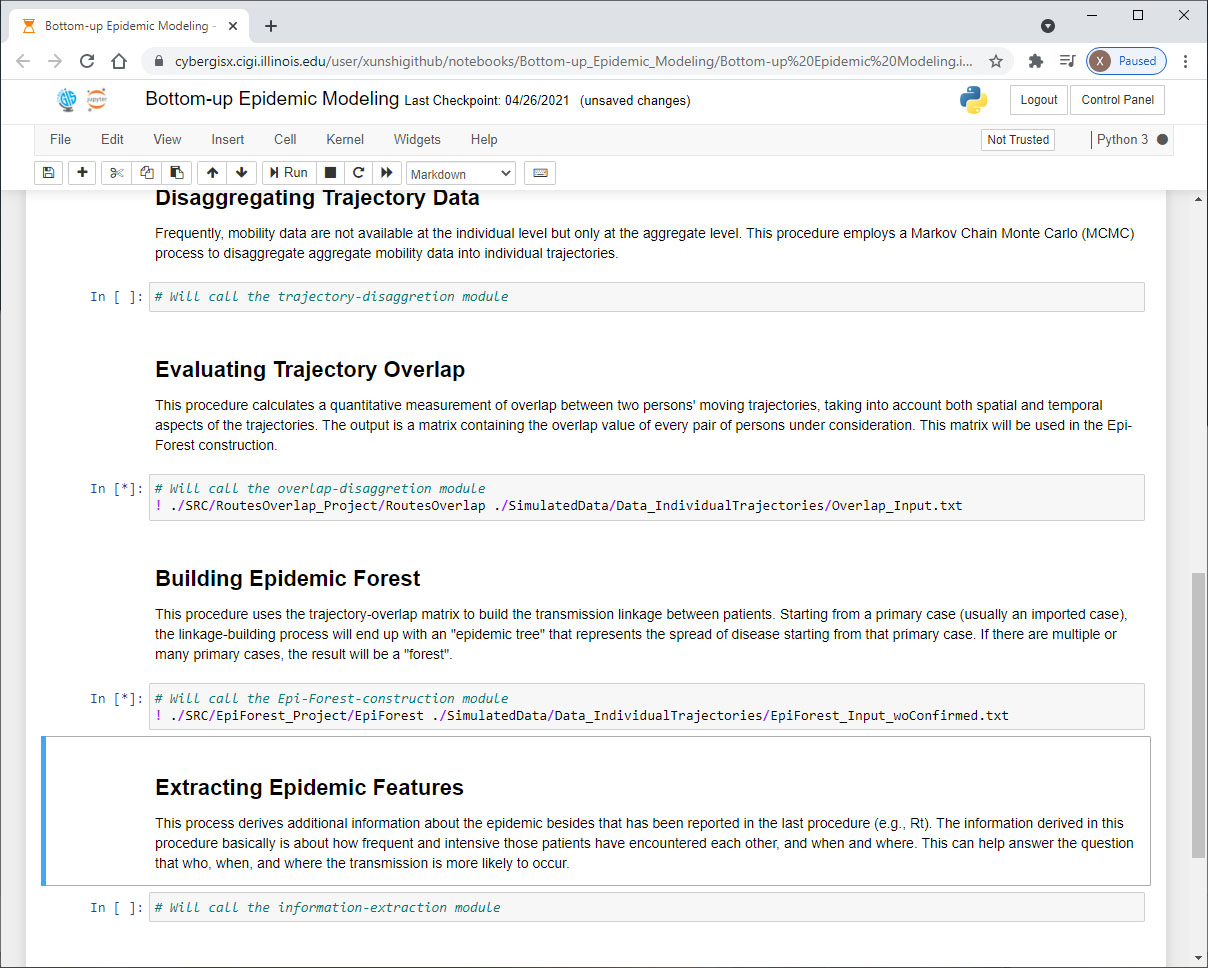
A Screenshots of Jupyter Notebook of the epi-forest modeling process on CyberGISX.
References:
Hakan Andersson and Tom Britton, 2000, Stochastic Epidemic Models and Their Statistical Analysis. Springer.
Li, M., Shi, X., Li, X., Ma, W., He, J., and Liu, T., 2019, Epidemic Forest: A Spatiotemporal Model of Communicable Diseases. Annals of the American Association of Geographers. 109(3): 812-836. DOI: 10.1080/24694452.2018.1511413.
Li, M., Shi, X., and Li, X., 2020, Integration of Spatialization and Individualization: The Future of Epidemic Modelling for Communicable Diseases, Annals of GIS, 26(3): 219-226. DOI: 10.1080/19475683.2020.1768438.
Shi, X. and Wang, S., 2015, Computational and Data Sciences for Health-GIS, Annals of GIS, 21(2): 111–118. DOI: 10.1080/19475683.2015.1027735.
